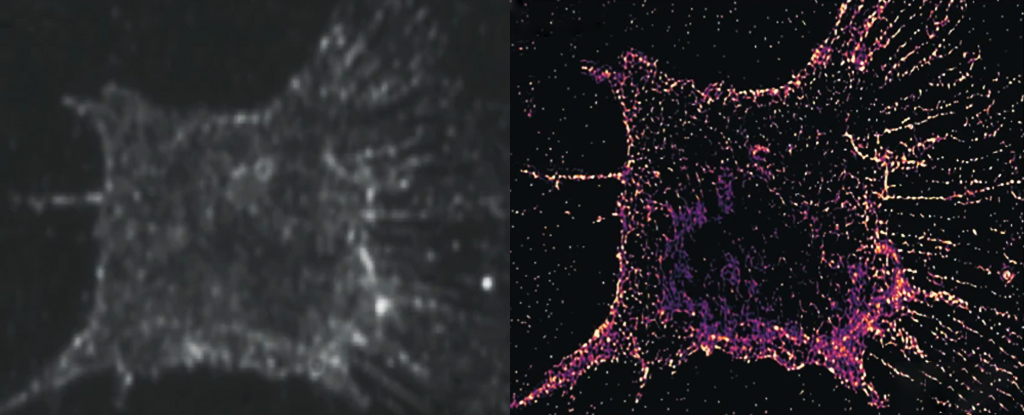
By replacing the fluorescent particles in the current imaging process with ones that scatter light instead, the researchers have revealed a whole new level of dazzling detail inside our living cells.
The innovative modification will allow scientists to directly observe molecular behavior over a much longer period, opening a window into pivotal biological processes such as cell division.
“A living cell is a place crowded with proteins here and there.” Explain University of Michigan Biomedical Engineer Guangjie Cui. “Our superior solution is very attractive to watch these dynamic activities.”
Super Resolution is a process for observing incredibly small biological structures. It uses a series of shots taken from groups of fluorescent particles that highlight specific areas of target tissue, eliminating the dulling effect of a flood of diffracted light.
The researchers behind its development won a Nobel Prize in 2014. As revolutionary as the process was, so was the ability of the fluorescent particles to be absorbed and then spit out The desired wavelength of light It wears out within tens of seconds, which excludes the assignment of longer duration operations.
So Cui and his colleagues developed a system to detect the scattering of light away from randomly distributed gold nanorods, a process that is not disrupted with repeated exposure to light. Although the gold markers are larger than the target structures, imaging multi-angle subsets of the bars and combining the images provide the same highly detailed resolution.
The resulting system allows for 250 hours of continuous observations with a resolution of just 100 atoms.
Cui and colleagues then examined the entire process of cell division using new PINE nanoscopy, revealing behavior not seen before. actin particlesdown to the individual molecule level.
Actin, the main component of the cell cytoskeletonProvides structural support to cells and helps facilitate movement within the cell. So these molecules in the form of branching filaments play a huge role in dividing the cell before separating it into two daughter cells.
Each copy of these cells inherits the same insides, from proteins to DNA, but exactly how this happens has long been a mystery due to the limitations of our optical technology.
By observing 904 actin filaments during the process of cell division, Cui and his team were able to see how the individual molecules behaved with each other. They found that when actin molecules were less bound together, they would expand in search of more bonds. As each actin reaches its neighbours, it approaches other actin molecules, causing their network to expand.
The researchers saw how these small-scale movements translated across a large-scale cellular display. Unexpectedly, when actin expands the cell in general, it actually contracts, whereas when actin contracts it expands. This seems paradoxical, so the researchers are keen to explore how this opposing movement occurs.
“We plan to use our method to study how other molecular building blocks are organized in tissues and organs,” said University of Michigan biomedical engineer Somin Lee. books for conversation.
“Our technology can help researchers visualize, and therefore better understand, how molecular defects in tissues and organs progress to disease.”
This research has been published in Nature Communications.

“Web maven. Infuriatingly humble beer geek. Bacon fanatic. Typical creator. Music expert.”





More Stories
Ryan Preece Penalized After Incident With Ty Gibbs at Texas Motor Speedway
Scientists Capture Elusive “Glowing Trees” Phenomenon During Thunderstorms
Satellite Images Reveal Newly Identified Driver of Greenland Glacier Melt